RESEARCH
Mechanistic Insights into Anti-Tumor Immune Responses
Cancer immunotherapy has become a central component of systemic treatment for various malignancies, including gastrointestinal cancers. However, in many cancer types, only approximately 10–20% of patients achieve durable clinical benefit from immunotherapy, and the mechanisms of resistance are considered to vary among individuals. Therefore, a deeper understanding of each patient’s tumor biology and appropriate treatment stratification are essential.
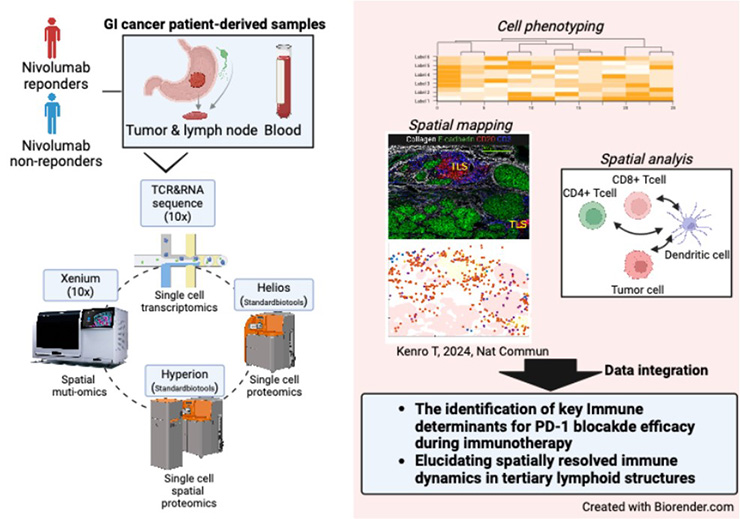
Our research group utilizes cutting-edge multi-omics technologies to analyze the characteristics and dynamics of immune cells, particularly T cells, at the single-cell level. In addition, advances in spatial biology have highlighted the importance of cellular localization and microenvironmental context in understanding disease mechanisms and identifying clinically relevant biomarkers.
Specifically, we perform spatial transcriptomics analysis using Xenium (10x Genomics) and T cell repertoire analysis using single-cell TCR and RNA sequencing (10x Genomics). Furthermore, to comprehensively profile immune responses in solid tumors and peripheral blood, we conduct spatial proteomics using the Hyperion Imaging System (Standard BioTools) and single-cell proteomic analysis using the Helios Mass Cytometer (Standard BioTools). Through these approaches, we have identified novel CD8⁺ T cell subsets associated with clinical responses to cancer immunotherapy (Kenro T, 2024, Nature Communications).
Our goal is to advance precision medicine by elucidating immune reprogramming at the single-cell level following immunotherapy and by uncovering mechanisms of therapeutic resistance. In parallel, we aim to define the functional organization and signaling networks within the tumor microenvironment, including tertiary lymphoid structures, to better understand anti-tumor immunity and improve patient outcomes.
Elucidation of Colorectal Cancer Initiation Mechanisms and Development of Novel Therapeutic Strategies
— Therapeutic Approaches Based on Cancer-Initiating (Stem) Cells, Epigenomic Alterations, and Clonal Hematopoiesis —
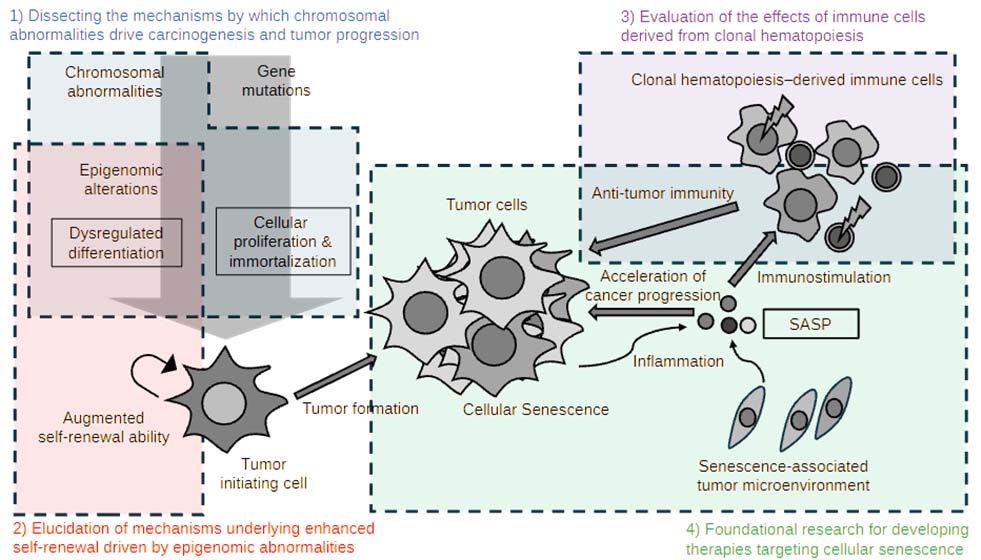
Cancer is a complex disease driven by multiple intertwined factors, and understanding the mechanisms underlying its initiation and progression is essential for the development of new therapeutic strategies. Recent studies have revealed that cancer-initiating (stem) cells, epigenomic modifications, and cellular senescence play critical roles in tumor initiation and progression. In our research, we aim to dissect the molecular mechanisms of colorectal carcinogenesis and to translate these findings into the development of novel therapeutic approaches.
Chromosomal Abnormalities, Epigenomic Dysregulation, and Carcinogenesis
Unlike normal cells, cancer cells frequently exhibit impaired celullar differentiation, which is often associated with dysregulation of epigenomic regulators. We investigate how specific epigenomic modifiers function in cancer-initiating (stem) cells and how their dysregulation leads to abnormal differentiation of cancer cells. In addition, chromosomal abnormalities represent one of the major drivers of carcinogenesis; however, their precise mechanistic contributions remain incompletely understood. In this study, we establish colorectal cancer models harboring defined chromosomal abnormalities and use these models to elucidate the processes of tumor initiation and progression. Through these efforts, we aim to develop novel therapeutic strategies targeting these genetic and epigenomic alterations.
Cellular Senescence, Clonal Hematopoiesis, and the Tumor Microenvironment
Recent studies suggest that clonal hematopoiesis may contribute to cancer development and progression. In this project, we perform detailed analyses of tumor microenvironmental changes induced by clonal hematopoiesis, with a particular focus on their impact on immune responses. Moreover, cellular senescence is known to promote tumor progression by inducing inflammation and immunosuppression within the tumor microenvironment. We therefore focus on the interplay between clonal hematopoiesis and senescent cells and aim to establish therapeutic strategies that target senescent cells in the tumor microenvironment.
Summary
We aim to elucidate the mechanisms of cancer initiation and progression from multiple perspectives, including cancer-initiating (stem) cells, epigenomic alterations, chromosomal abnormalities, clonal hematopoiesis, and cellular senescence, and to develop novel therapeutic approaches based on these insights. In particular, we pursue translational research bridging basic and clinical studies, such as investigations into differentiation defects of colorectal cancer stem cells, the relationship between clonal hematopoiesis and the tumor microenvironment, and the development of senolytic-based cancer therapies. Progress in these studies is expected to enable more effective treatments for cancer patients and to represent an important step toward overcoming cancer.
Identification of Predictive Factors for the Efficacy of Immune Checkpoint Inhibitors
In recent years, advances in cancer molecular biology have revealed that specific molecular targeted therapies exhibit high efficacy against cancers harboring so-called driver mutations, such as human epidermal growth factor receptor (EGFR) mutations and ALK fusion genes. It has also been elucidated that cancer cells can evade immune surveillance mechanisms through immune checkpoint molecules such as PD-1 and PD-L1. Consequently, immune checkpoint inhibitor (ICI) targeting these pathways has been introduced for various cancer types.
Several tumor-related factors have been reported as predictors of ICI efficacy, including PD-L1 expression in tumor tissues, deficiency in DNA mismatch repair (MMR) and microsatellite instability (MSI), as well as tumor mutational burden (TMB). However, no established biomarkers have been identified among host (immune system) factors. Moreover, the effects of ICI therapy on immune cells other than T cells within the tumor microenvironment, as well as the subsequent systemic immune responses, remain insufficiently elucidated.
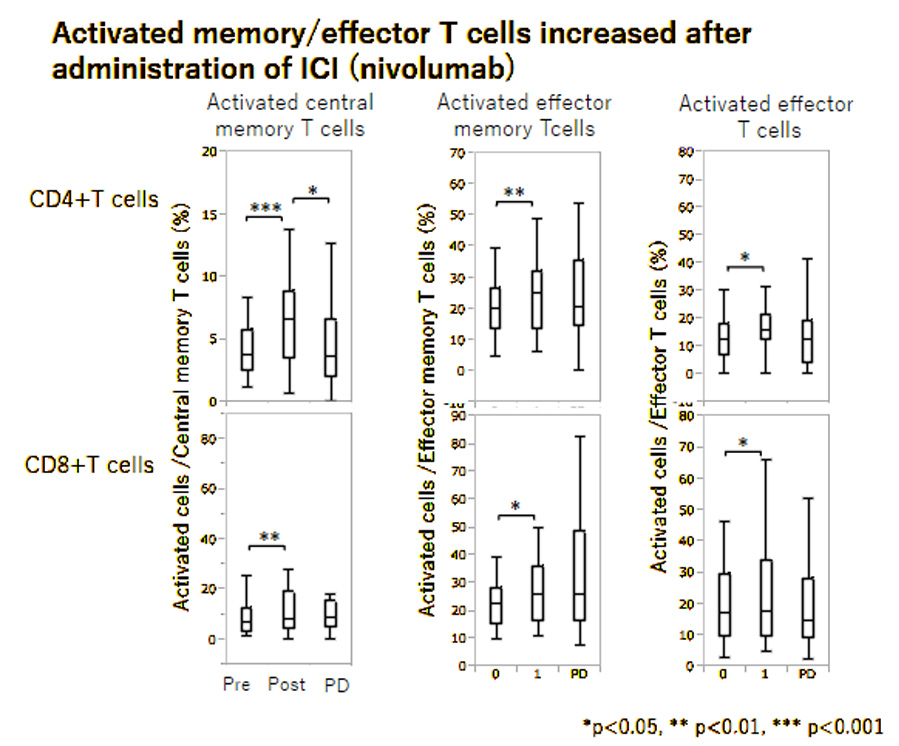
In our department, we are analyzing immune cell subsets and profiling cytokines using blood and tissue samples from patients treated with ICIs, aiming to identify predictive factors for therapeutic efficacy and resistance, as well as specific immune cell subsets involved in tumor immunity. We are also investigating the functions of these cells. Through this research, we hope to contribute to the development of more precise cancer immunotherapies.
Treatment and Blood Sampling
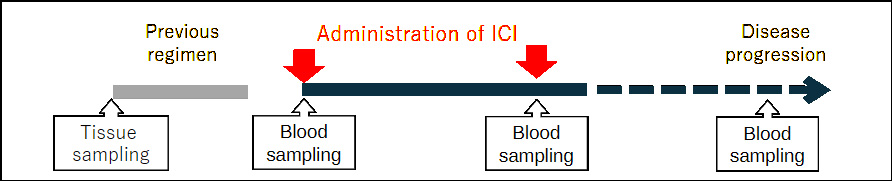
Analysis using Flow Cytometry
- Peripheral blood mononuclear cells (PBMCs):
Analysis of immune cell subsets based on surface antigen expression - Serum: Quantification of cytokines
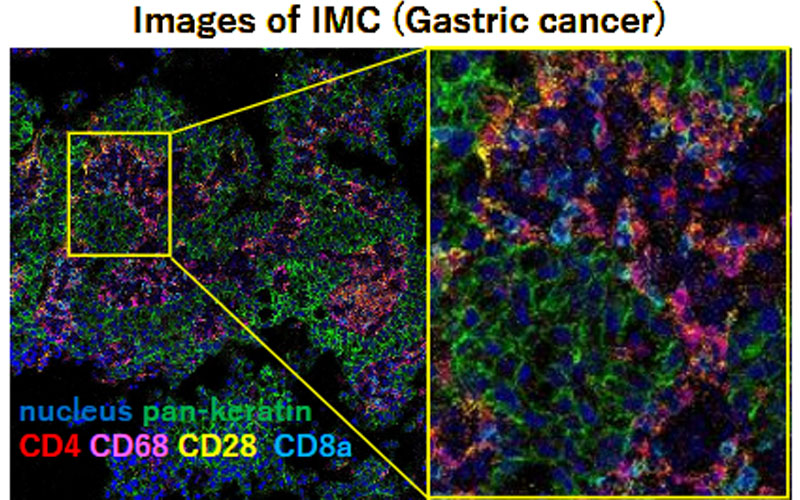
Analysis using Imaging Mass Cytometry (IMC)
- Analysis of immune cell subsets in the tumor microenvironment
Origins and Control of Spontaneous Mutations— DNA Damage, Repair, and Mutagenesis in Somatic and Germ Cells–
Understanding how genetic mutations arise and how organisms suppress them is essential for uncovering the fundamental principles of life. Mutations in somatic cells can lead to cancer and aging, while those in germ cells may result in infertility, miscarriage, or hereditary disorders. More broadly, over long evolutionary timescales, mutations generate biological diversity and serve as drive evolution. Thus, elucidating the mechanisms by which spontaneous mutations occur and how they are prevented is key to understanding the essence of life.
Reactive oxygen species produced during cellular energy metabolism induce DNA damage and contribute to spontaneous mutations. Yet many questions remain: How does oxidative DNA damage drive mutagenesis and cancer development? How does germline DNA damage affect the genomes of offspring and later generations? Ionizing radiation and chemical agents also generate diverse types of DNA lesions, but their impacts on genetic information and disease risk are not fully understood.
To address these issues, we use DNA repair–deficient knockout mice to experimentally evaluate the origins and consequences of DNA damage. By investigating both damage formation and repair mechanisms, we aim to deepen our understanding of how living organisms maintain the integrity of their genomes. These studies not only advance our knowledge of cancer and hereditary disease mechanisms but also shed light on the evolutionary processes shaped by mutations.
Major Research Projects
- Somatic Mutation Project: Oxidative DNA damage and gastrointestinal carcinogenesis
- Germline Mutation Project: Identifying factors that determine mutation rates and mutation spectra
- Oocyte DNA Repair Project: Oocyte-driven repair of sperm DNA damage and its role in embryonic development
- Biological Effects of Radiation: Transgenerational mutation effects; internal ¹³⁷Cs exposure in multi-generational mice
- Education-Related Projects: Educational content for genetics and radiation-effect literacy
Germline mutation project
How do de novo mutations occur during the life cycle of germ cells? In this project, we conduct experimental analyses using mouse strains deficient in DNA repair mechanisms to elucidate the types of DNA damage that primarily cause mutations, the timing and extent of their accumulation, and the DNA repair pathways that suppress them. Through this research, we aim to deepen our understanding of the mechanisms that maintain genome stability in germ cells and gain new insights into the regulation of genetic mutations transmitted to offspring. Additionally, utilizing these experimental systems, we analyze the effects of various environmental factors, such as radiation and chemical substances.
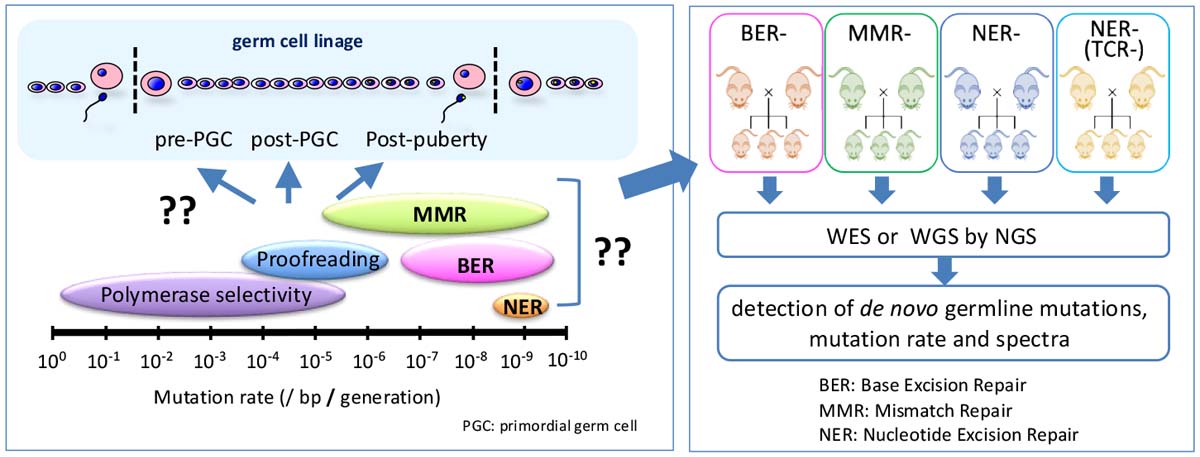
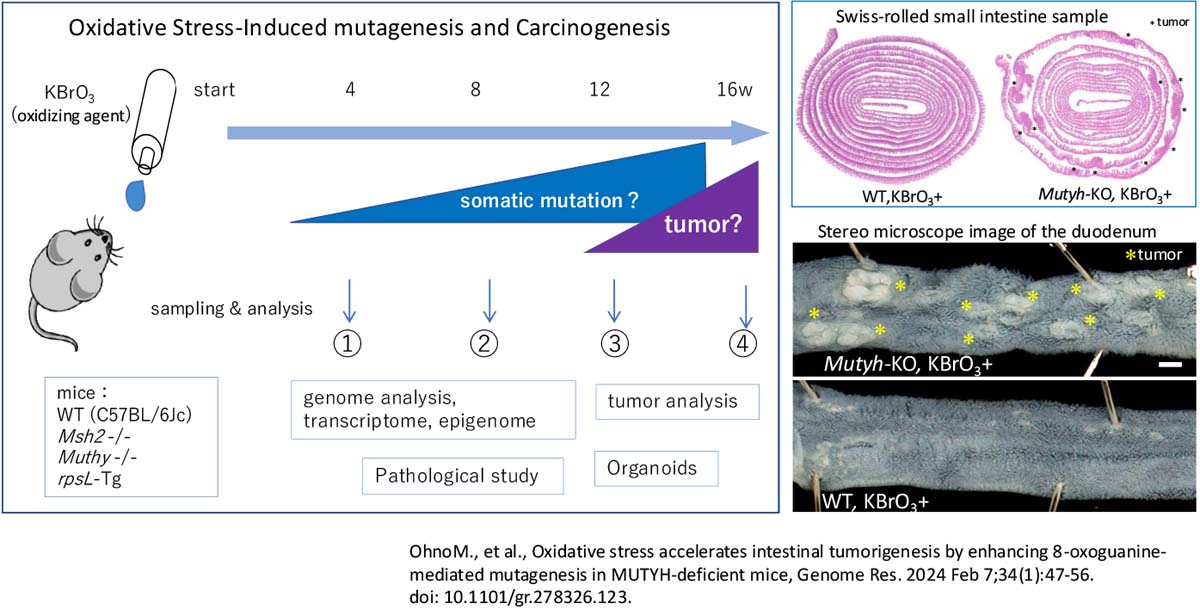
Somatic mutation project
This project aims to elucidate the mechanisms of carcinogenesis caused by oxidative DNA damage and defects in its repair system. Using knockout mice for genes such as Mutyh and Msh2, which are known as causative genes of hereditary colorectal cancer, we are investigating the relationship between oxidative stress, mutagenesis, and carcinogenesis.
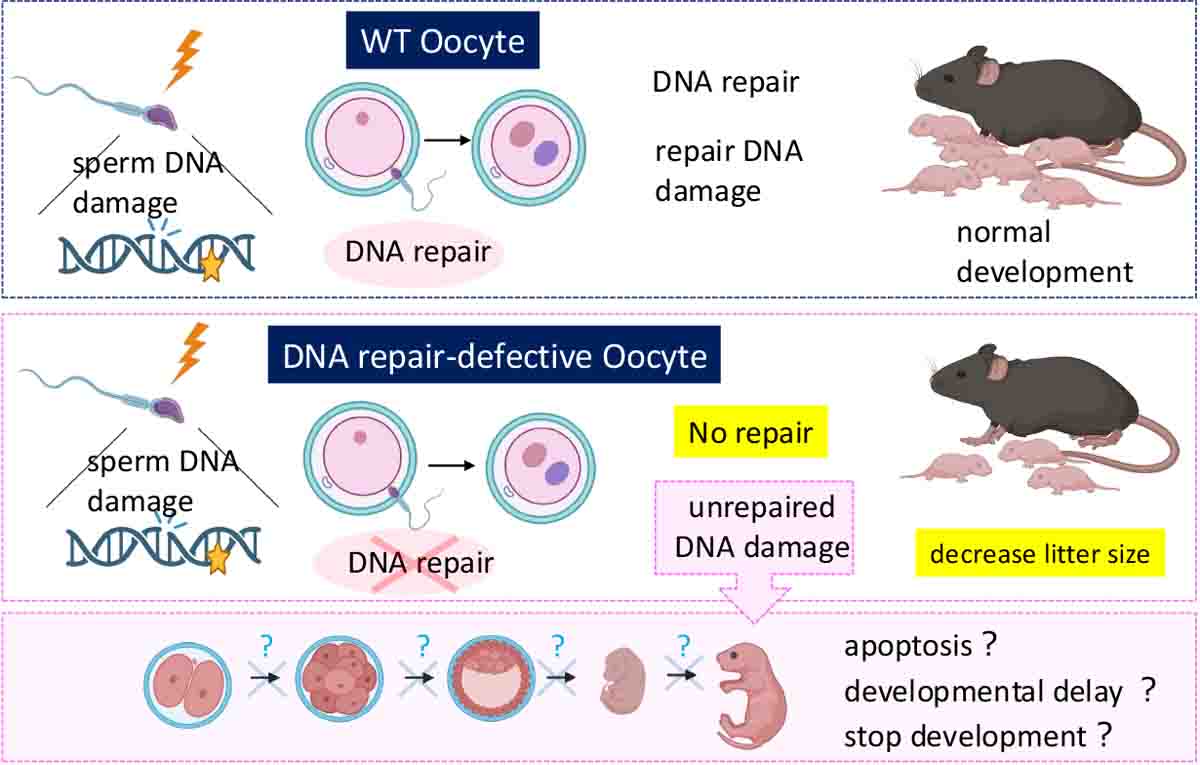
Oocyte DNA Repair Project : Unveiling Its Role in Embryonic Genome Integrity
Sperm DNA damage is recognized as a risk factor for infertility and miscarriage, while the oocyte’s DNA repair mechanisms are suggested to contribute to normal embryonic development. This study aims to elucidate the underlying mechanisms by using DNA repair-defective mice.